
Derek Dee
Assistant Professor
I am a food scientist and biophysicist, having focused on the folding mechanisms of aspartic proteases, the misfolding and aggreagation of prion protein at the single molecule level, and the conversion of food related proteins into nanofibrils.
Office FNH 240
Office phone 604-822-4489
Email
Graduate Students

Drew Sanders
Ph.D. candidate, co-supervised with prof. Rickey Yada.
I am using optical tweezers to investigate pepsin and pepsinogen folding at the single molecule level. These proteins are an excellent model for studying protein (kinetic) stability, which remains poorly understood, and characterizing how evolution sculpts protein folding energy landscapes. Better understanding these basic properties may have utility in engineering novel, industrially-relevant enzymes for stable catalysis under a wider range of conditions and in targeting pepsin-like proteases involved in several diseases.

Leslie Huynh
Master’s student, co-supervised with prof. Rickey Yada.
I joined the Dee Lab in September 2023 after completing my Bachelor’s degree in Food Science at the University of Guelph. My research focuses on recombinant expression and characterization of pepsin-like proteases from cold-adapted yeast. We want to better understand how extremophilic enzymes have adapted to fold and function in harsh environments. Besides protein engineering, this research can be useful in applications which require lower temperatures, such as cold-wash detergents, leather manufacturing and cold-ripening of cheeses.
Email

Amalia Diaz de Leon Derby
Master’s student
I completed my bachelor’s in Engineering in biotechnology at Tecnológico de Monterrey and joined the Dee Lab in January 2025. My research focuses on the recombinant expression and characterization of CsgA, the main structural protein of curli amyloid fibers produced by E. coli. These functional amyloids play a critical role in biofilm formation. I aim to investigate how specific regions contribute to fibril assembly and stability. In my free time I enjoy going for runs and being outside. I also enjoy painting and cooking.
Email

Research Associates
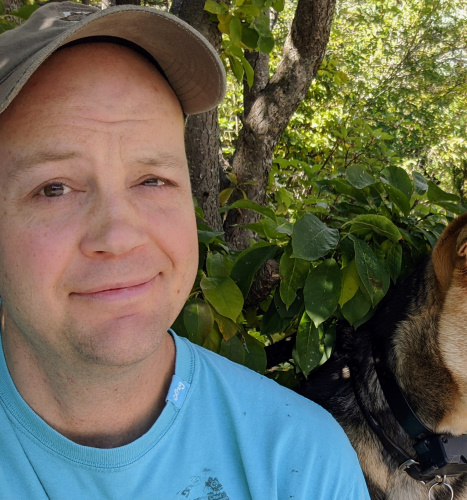

Daniel Foster
Ph.D. Physics. (Focus areas: biophysics and optics)
I joined Derek’s lab in the fall of 2021, to help with the single molecule biophysics work using optical tweezers for force spectroscopy. During my graduate studies at the University of Alberta, I was part of a team who built three optical tweezers instruments in Michael Woodside’s lab. Here at UBC we have the commercial Lumicks’ C-trap with which we explore protein folding.
My graduate research focused on both optical tweezers construction (optics, alignments, programming, CAD) and advancing measurement capabilities, while studying the folding of nucleic acids: both the DNA hairpin model systems, and RNA regulatory elements such as riboswitches and frame shifting pseudoknots. My post doctoral research involved the prototyping of microfluidic devices (from the cleanroom to the wet bench) and the (stop/slit) flow lithography of functional hydrogels (fluophores and functional proteins) in Dae Kun Hwang’s lab in Toronto.
Email
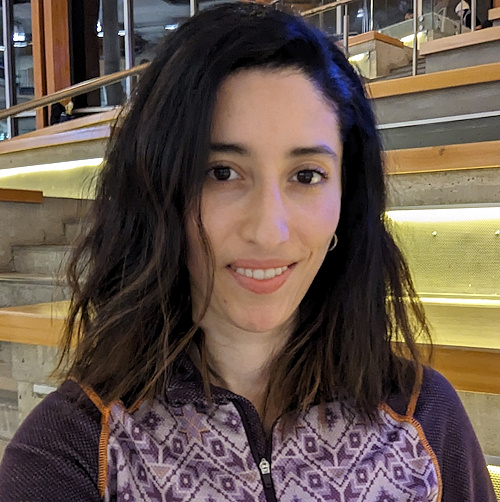

Aicha Asma Houfani
Ph.D. Microbiology.
I joined Derek’s lab to use advanced proteomics tools to characterize seed-storage proteins from legumes, focusing on pea proteins initially, to understand how protein structural changes contribute to quality attributes of pea protein used in food.
With a microbiology background from bachelor’s to PhD, I have spent the past few years as a postdoc at UBC, diving into three major research projects. Along the way, I have gained expertise in mass spectrometry-based proteomics and protein biochemistry through collaborations with local industries via Mitacs programs and BC Cancer at Leonard Foster Lab’s to develop and apply (phospho)proteomics for studying the mechanism of action of a small molecule drug and combination therapeutics, as well as developing proteomics for protein corona characterization with lipid nanoparticles. During my PDF, I was working with the NRC and Derek Dee’s Lab on a cutting-edge sustainable protein production project focused on legume proteomics. My current focus is back in microbiology to explore plant-specific inserts (PSI) from Solanum tuberosum and their antifungal properties, working with Dr. Lennie Cheung and Prof. Rickey Yada. This takes the PSI from recombinant protein expression experiments to direct introduction into fungal assays.
Email
Summer students in 2025

Rachel Rosenberg (LFS senior undergraduate honours thesis)
I am currently doing my undergraduate degree in food science at UBC. At the Dee lab this summer I will be working on and refining skills pertaining to recombinant protein expression and purification. I am passionate about creating sustainable food systems via precision fermentation/recombinant protein expression to create alternative protein sources.


Perry Sandhu (NSERC USRA)
I am a biochemistry undergraduate student at UBC. At the Dee lab I help study protein folding on a single-molecule scale. This summer I will work on recombinant protein production, synthesizing DNA handles, and force spectroscopy on these protein-DNA constructs using optical tweezers.


Annika Dzieia (Mitacs Globalink)
working on pea protein fibrils


Julia Gutierrez Castillejos (Mitacs Globalink)
working on CsgA, _B, _C, and the purification of these proteins.

Lab alumni:
- Hrishika Dekate (MITACS Globalink intern 2024)
- Lanfang ‘Charlotte’ Shi (M.Sc. Dec 2023), Beverage technologist at the Mark Antony Group
- Alexandra Reith (Mitacs Globallink summer student 2023) Hochschule Darmstadt, Germany
- Jogindh Sivakumar Suganthi (Mitacs Globallink summer student 2023) IISER Pune, India to Max-Planck-Institute for Terrestrial Microbiology
- Fan Bu (2021-2022) transferred to U. Minn. Pharmacology*
- Yuran Zhang (M.Sc. Dec 2022), Biochemistry Research Scientist at CarboNet
- Vivian Zhang -> Wageningen U (wur.nl) (NSERC summer student 2022)
- Yu He (Mitacs Worklearn summer student 2022), UBC
- Prerana Balasubramanian (Mitacs Globalink summer intern 2022), at UMass Amherst
- Sara Zamani (MSc, Dec 2021)
- Sneha Kelkar (BSc)
- Alyssa Robertson (MSc, UGA)
- Lida Rahimi Araghi (MSc, UGA)
- Ana Jaworski (MSc, UGA)
Former visiting Dekaban scolars:
- Katarzyna ‘Kasia’ Skrzypczak (Lublin, Poland) 2024
- Bartosz G. Sołowiej, Vice-Rector (Lublin, Poland)2022